Can windows system install hmmer software
hmmer download and installation
For Mac OS/X, Linux, UNIX systems, compile and install from source code:
% wget ftp://selab.janelia.org/pub/software/hmmer3/3.0/hmmer-3.0.tar.gz % tar zxf hmmer-3.0.tar.gz % cd hmmer-3.0 % ./configure % make % make check
For Windows systems, download the binary compressed package directly, decompress it and use it.
Programs included in hmmer
phmmer: Similar to Blastp, uses a protein sequence to search the protein sequence library;
>phmmer tutorial/HBB HUMAN uniprot sprot.fa
jackhmmer: Similar to psiBlast, protein sequences iteratively search protein sequence libraries;
>jackhmmer tutorial/HBB HUMAN uniprot sprot.fa
hmmbuild: Build HMM model using multiple alignment sequences;
hmmsearch: Use HMM model to search sequence library;
hmmscan: Use sequence to search HMM library;
hmmalign: Use HMM as a clue to construct multiple alignment sequences;
>hmmalign globins4.hmm tutorial/globins45.fa
hmmconvert: Convert HMM format
hmmemit: Obtain a pattern sequence from the HMM model;
hmmfetch: Retrieve an HMM model from the HMM library by name or acceptance number;
hmmpress: Format the HMM database to facilitate hmmscan search and use;
hmmstat: Display statistical information of HMM database;
Search sequence database using HMM model
Use hmmbuild to build the HMM model, input a multiple alignment sequence file in Stockholm format or FASTA format (such as: tutorial/globins4.sto), the command is as follows:
>hmmbuild globins4.hmm tutorial/globins4.sto
globins4.hmm is the output HMM model
Use hmmsearch to search the protein sequence database. The protein sequence database is in FASTA format. The command is as follows:
>hmmsearch globins4.hmm uniprot sprot.fasta >globins4.out
globins4.out is the output result file, as follows:
*The example uses the example in the official tutorial
Search the HMM database using protein sequences
Build an HMM database. The HMM database is a file containing multiple HMM models. It can be downloaded from Pfam, SMART, and TIGRFams, or it can be built by yourself from multiple alignment sequences, such as:
>hmmbuild globins4.hmm tutorial/globins4.sto
>hmmbuild fn3.hmm tutorial/fn3.sto
>hmmbuild Pkinase.hmm tutorial/Pkinase.sto
>cat globins4.hmm fn3.hmm Pkinase.hmm >minifam
Use hmmpress to format the database, including compression and index creation. The command is as follows:
>hmmpress minifam
This step can be completed quickly, and the output content is as follows:
Working… done.
Pressed and indexed 3 HMMs (3 names and 2 accessions).
Models pressed into binary file: minifam.h3m
SSI index for binary model file: minifam.h3i
Profiles (MSV part) pressed into: minifam.h3f
Profiles (remainder) pressed into: minifam.h3p
Use hmmscan to search the HMM database, the command is as follows:
>hmmscan minifam tutorial/7LESS_DROME
How does hmmer software convert fasta format files to sto format
I also encountered this problem. I searched online for a long time and couldn't find a suitable solution, so I wrote one myself. The code is as follows
import glob # They are all things from the standard library
import os
# Put the fasta file (compared) you want to build hmm in the same folder as this program, and then run this program to run hmmbuild
os.chdir(os.path.dirname(__file__))
fs = glob.glob('*.fasta') # Get each fasta file. If your fasta file has a suffix other than .fasta, you can change it here, or directly change it to '*.fa*'
for f in fs:
hmm = os.path.splitext(f)[0] '.hmm'
stockholm = os.path.splitext(f)[0] '.sto'
with open(f, 'r') as fhandle: # This is used to read fasta files and save all fasta files to the list
fastas = ['>' tmp.replace('\n', '\r', 1).replace('\n', '').replace('\r', '\n') for tmp in tuple(filter(None, (fhandle.read().split('>'))))]
for i in range(len(fastas)):
fastas[i] = fastas[i].split('\n')
fastas[i][0] = fastas[i][0].split()[0][1:10]
tmp = []
for j in range(len(fastas[i][1]) // 80 1):
tmp.append(fastas[i][1][80 * j : 80 * j 80])
fastas[i][1] = tmp
with open(stockholm, 'w') as out: # The sto file is written here
out.write('# STOCKHOLM 1.0\n\n')
for j in range(len(fastas[0][1]) - 1):
for i in range(len(fastas)):
out.write('% -12s%s\n' % (fastas[i][0], fastas[i][1][j]))
out.write('\n')
for i in range(len(fastas)):
out.write('% -12s%s\n' % (fastas[i][0], fastas[i][1][-1]))
out.write('//')
os.system('hmmbuild --amino %s %s' % (hmm, stockholm)) # hmmbuild is running here, you can modify the parameters inside
How to teach yourself bioinformatics
1. Start with existing bioinformatics tools. Be familiar with how to use existing software, network servers, databases, etc. to serve biological research. Don’t repeat work. Don’t develop your own if you can use ready-made ones.
2. Familiar with command line operating systems, such as DOS and Linux, you can write simple shells; then you can install command line-level programs and run some regular processes. Learning how to find and install software is the most important and fundamental skill. In fact, many problems can be easily solved if you find the right software package.
3. Be familiar with a simple scripting language. I personally recommend using python. For specific reasons, please see my post. Small scripts are very useful when there are no ready-made tools, or when data format conversion is required. General applications do not need to write too much code by themselves. We must believe that other experts may have encountered the problems we usually encounter, so there are a large number of toolkits on the Internet. As for more programming languages, one can master everything, R, perl, etc. are all similar.
4. Be familiar with the knowledge of simple algorithms and data structures, so that you can understand the internal mechanisms of many programs, and then know their advantages and disadvantages, which will also be helpful for writing your own programs. If you have the energy, then study statistics, machine learning, etc. .
5. Expand, research, analyze, and develop within your own biological field.
The above is the detailed content of Can I install HMMER software on Windows systems?. For more information, please follow other related articles on the PHP Chinese website!
 Acer PD163Q Dual Portable Monitor Review: I Really Wanted to Love ThisMar 18, 2025 am 03:04 AM
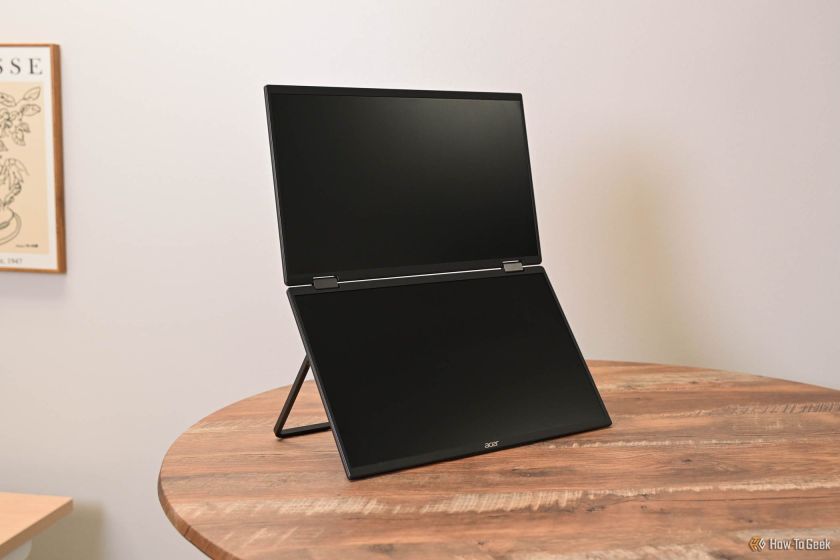
Acer PD163Q Dual Portable Monitor Review: I Really Wanted to Love ThisMar 18, 2025 am 03:04 AMThe Acer PD163Q Dual Portable Monitor: A Connectivity Nightmare I had high hopes for the Acer PD163Q. The concept of dual portable displays, conveniently connecting via a single cable, was incredibly appealing. Unfortunately, this alluring idea quic
 win11 activation key permanent 2025Mar 18, 2025 pm 05:57 PM
win11 activation key permanent 2025Mar 18, 2025 pm 05:57 PMArticle discusses sources for a permanent Windows 11 key valid until 2025, legal issues, and risks of using unofficial keys. Advises caution and legality.
 win11 activation key permanent 2024Mar 18, 2025 pm 05:56 PM
win11 activation key permanent 2024Mar 18, 2025 pm 05:56 PMArticle discusses reliable sources for permanent Windows 11 activation keys in 2024, legal implications of third-party keys, and risks of using unofficial keys.
 Top 3 Windows 11 Gaming Features That Outshine Windows 10Mar 16, 2025 am 12:17 AM
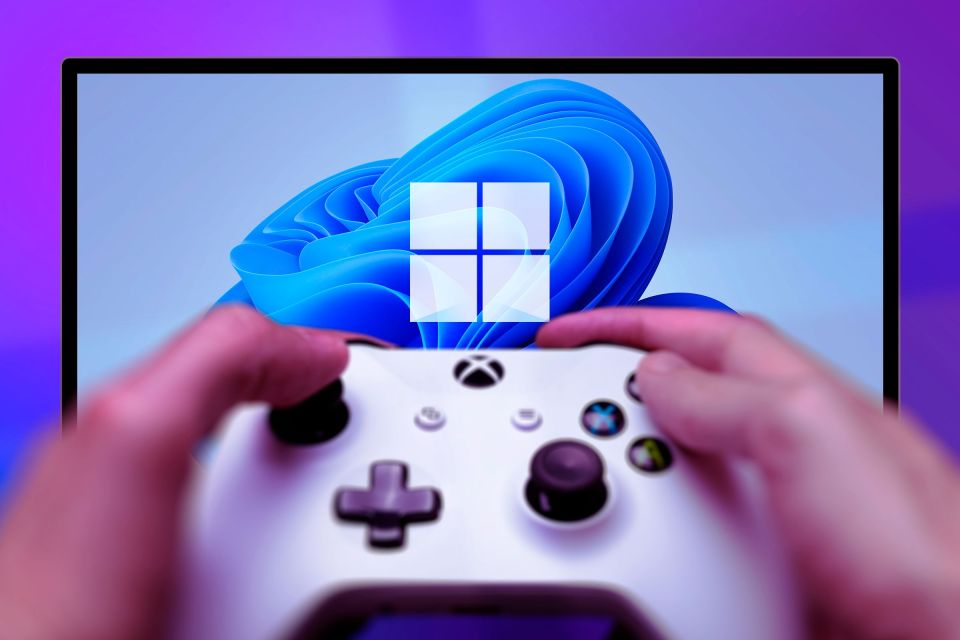
Top 3 Windows 11 Gaming Features That Outshine Windows 10Mar 16, 2025 am 12:17 AMUpgrade to Windows 11: Enhance Your PC Gaming Experience Windows 11 offers exciting new gaming features that significantly improve your PC gaming experience. This upgrade is worth considering for any PC gamer moving from Windows 10. Auto HDR: Eleva
 Mozilla Thunderbird 136 Is Here, Switching to Monthly Updates by DefaultMar 07, 2025 am 01:19 AM
Mozilla Thunderbird 136 Is Here, Switching to Monthly Updates by DefaultMar 07, 2025 am 01:19 AMFirefox 136 and Thunderbird 136: Enhanced Security and Performance The latest releases of Firefox and Thunderbird bring significant improvements in video playback smoothness, browsing security, and overall user experience. Let's delve into the key u
 The Best Monitor Light Bars of 2025Mar 08, 2025 am 03:02 AM
The Best Monitor Light Bars of 2025Mar 08, 2025 am 03:02 AMReduce eye strain and brighten your workspace with a monitor light bar! These handy gadgets adjust brightness and color temperature, some even offering auto-dimming. This updated review (03/04/2025) highlights top picks across various needs. BenQ
 How to Create a Dynamic Table of Contents in ExcelMar 24, 2025 am 08:01 AM
How to Create a Dynamic Table of Contents in ExcelMar 24, 2025 am 08:01 AMA table of contents is a total game-changer when working with large files – it keeps everything organized and easy to navigate. Unfortunately, unlike Word, Microsoft Excel doesn’t have a simple “Table of Contents” button that adds t
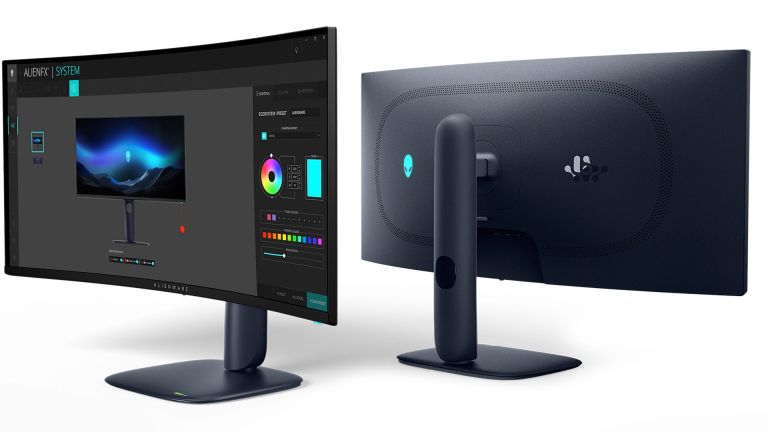 This Wild Ultra-Wide Alienware Monitor is $300 Off TodayMar 13, 2025 pm 12:21 PM
This Wild Ultra-Wide Alienware Monitor is $300 Off TodayMar 13, 2025 pm 12:21 PMAlienware AW3225QF: The best curved 4K display, is it worth buying? The Alienware AW3225QF is known as the best curved 4K display, and its powerful performance is unquestionable. The fast response time, stunning HDR effects and unlimited contrast, coupled with excellent color performance, are the advantages of this monitor. Although it is mainly aimed at gamers, if you can accept the shortcomings of OLED, it is also suitable for office workers who pursue high efficiency. Widescreen monitors are not only loved by gamers, but also favored by users who value productivity improvement. They are great for work and enhance anyone’s desktop experience. This Alienware monitor is usually expensive, but is currently enjoying it


Hot AI Tools
Undresser.AI Undress
AI-powered app for creating realistic nude photos
AI Clothes Remover
Online AI tool for removing clothes from photos.

Undress AI Tool
Undress images for free

Clothoff.io
AI clothes remover

AI Hentai Generator
Generate AI Hentai for free.

Hot Article

Hot Tools

Dreamweaver CS6
Visual web development tools

Zend Studio 13.0.1
Powerful PHP integrated development environment

Safe Exam Browser
Safe Exam Browser is a secure browser environment for taking online exams securely. This software turns any computer into a secure workstation. It controls access to any utility and prevents students from using unauthorized resources.

SublimeText3 Mac version
God-level code editing software (SublimeText3)

Atom editor mac version download
The most popular open source editor